Our citizen science project, Rodent Little Brother, which has been developed to help scientists identify early disease characteristics through the study of mouse behaviour in the home-cage environment, has won an Understanding Animal Research Openness Award!
Rodent Little Brother engages the public in analysing mouse behaviour and activity through watching video clips of laboratory mice housed in their home-cage and annotating each clip. The data the citizen groups produce will be used to develop a machine learning algorithm that will allow mouse interactions and behaviour to be monitored in the home-cage environment and analysed automatically, without the need for human intervention. The analysis of mouse behaviour helps researchers identify the potential role of genes in disease and possible routes for therapy.
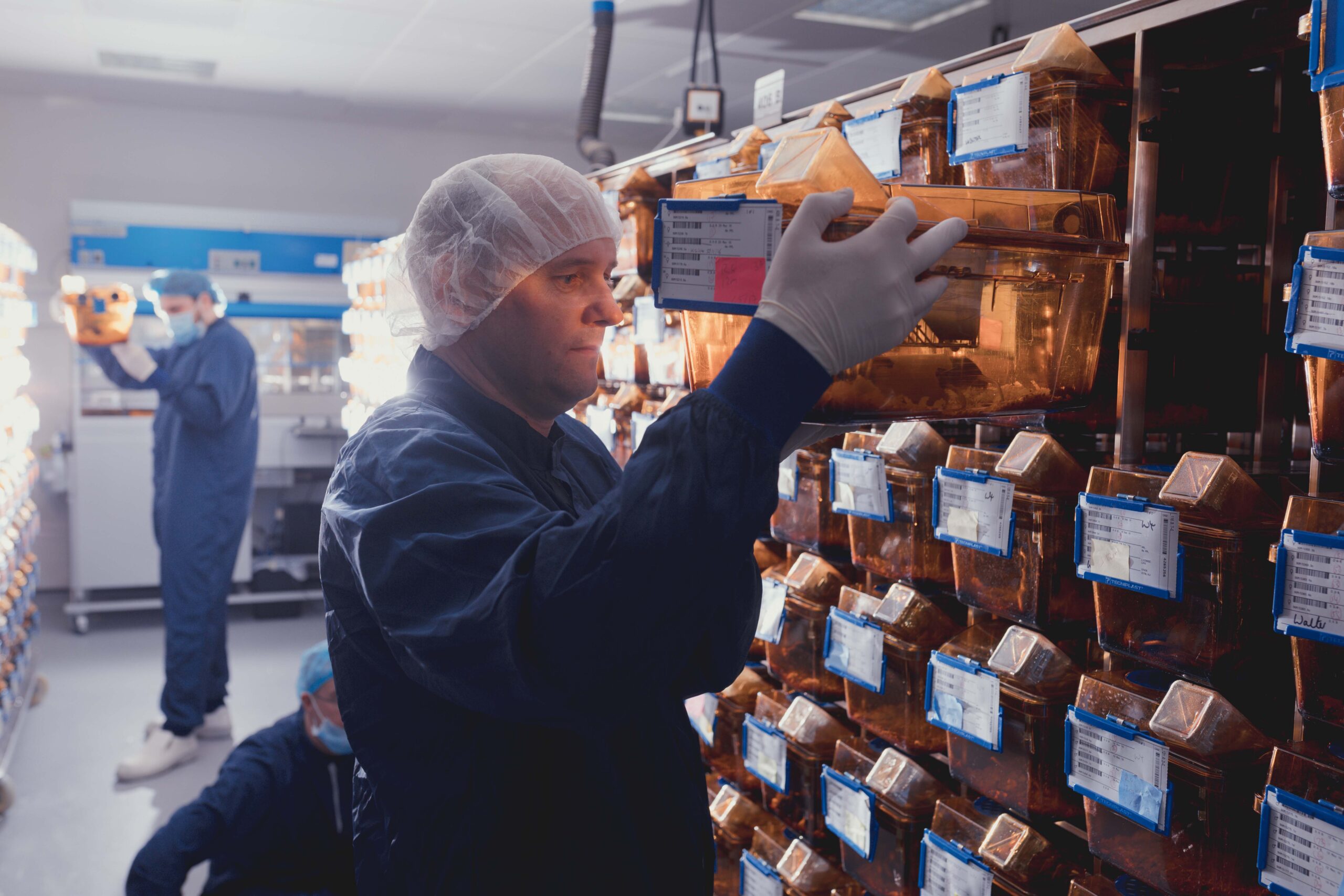
Mice are used widely in research due to their similarity in both their DNA and organ function to humans. Researchers commonly study genetically altered mouse models, by modifying a particular gene or set of genes in the mouse genome. They can then identify how that may lead to disease by studying the mouse’s behaviour, interactions with other mice and daily activities, such as eating and sleeping, to observe any implications of the genetic change. However, tests often require mice to be removed from their cage, isolated from social groups for the test duration, and are regularly carried out during hours of light, despite mice being primarily active during the hours of darkness. This might mean some behaviours that would normally be displayed go unrecorded due to mice being in unusual settings. To improve the science and obtain more accurate recordings of data, it would be beneficial to study the mouse in its home-cage, where normal interactions can be observed.
As such, researchers at the Mary Lyon Centre worked with Actual Analytics in Edinburgh to develop the Home Cage Analysis system (HCA) as part of the NC3Rs CRACK IT challenge. The HCA allows for mouse movement to be tracked in an unobtrusive and continuous manner via video recording and infrared LED lighting. These video recordings allow researchers to study behaviours in social settings across various time periods. However, analysing and reporting data from these videos is extremely time-consuming. The Rodent Little Brother project nicely combines the need for analysing hundreds of minutes of clips with the benefit of improving mouse welfare, by monitoring mice in their social groups within the home-cage settings.
This project has been highly beneficial in allowing our institute and our researchers to create a dialogue with audiences globally, about the nature and the need for laboratory animal science. There is a list of activities for the users to choose from and a discussion board for the participants to share their experiences and thoughts. In addition, a talk board is available which is aimed solely at the research team where the participants may ask questions of the team, directly. Throughout the project a true two-way conversation has developed between the public and scientists. This has created both an open and honest discussion around the essential role of mice in medical research as well as an understanding of the thoughts and ideas of the public in this research.
We are delighted to be awarded this prize. This project is a testament to our continued commitment to engage in open conversations with a wide variety of audiences, with varied views and backgrounds in the context of animal research and the wider scientific field. We are particularly thankful for the support we have received from the MRC, NC3Rs and Zooniverse for making this project possible.
Dr. Rasneer Sonia Bains, Project lead on Rodent Little Brother Project